|
55 |
7BQY |
The Crystal Structure of COVID-19 Main Protease in Complex with an Inhibitor N3 at 1.7 Å [Structure of Mpro from COVID-19 virus and discovery of its inhibitors: 6LU7と同一論文] |
04/22/2020 |
|
54 |
6WEY |
High-resolution structure of the SARS-CoV-2 NSP3 Macro X domain |
04/29/2020 |
|
53 |
6WLC |
Crystal Structure of NSP15 Endoribonuclease from SARS CoV-2 in the Complex with Uridine-5'-Monophosphate (関連 ID 6X1B 6VWW 6W01 6WLC 6WXC) [Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2.] |
04/29/2020 |
|
52 |
6YHU |
Crystal structure of the nsp7-nsp8 complex of SARS-CoV-2 |
04/29/2020 |
|
51 |
6YM0 |
Crystal structure of the SARS-CoV-2 receptor binding domain in complex with CR3022 Fab (crystal form 1) [Potent antibody binding to an unexpected highly conserved cryptic epitope of the SARS-CoV-2 Spike, to be published] - 6YLA (released 2020-04-15)関連 |
04/29/2020 |
|
50 |
6WKP |
Crystal structure of RNA-binding domain of nucleocapsid phosphoprotein from SARS CoV-2, monoclinic crystal form |
04/29/2020 |
|
49 |
6W37 |
SARS-CoV-2 ORF7a encoded accessory protein accessory protein |
04/29/2020 |
|
48 |
6LZE |
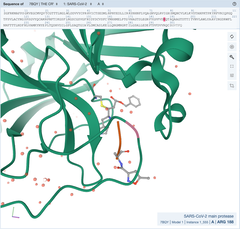
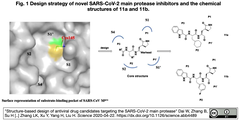
SARS-CoV-2 Mpro in complex with 11a [Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease] |
04/29/2020 |
|
47 |
6M0K |
SARS-CoV-2 Mpro in complex with 11b [Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease] |
04/29/2020 |
|
46 |
6WJI |
2.05 Angstrom Resolution Crystal Structure of C-terminal Dimerization Domain of Nucleocapsid Phosphoprotein from SARS-CoV-2 |
04/22/2020 |
|
45 |
6WIQ |
Crystal structure of the co-factor complex of NSP7 and the C-terminal domain of NSP8 from SARS CoV-2 |
04/22/2020 |
|
44 |
7BV1 |
Cryo-EM structure of the apo nsp12-nsp7-nsp8 complex |
04/22/2020 |
|
43 |
7BV2 |
The nsp12-nsp7-nsp8 complex bound to the template-primer RNA and triphosphate form of Remdesivir(RTP) |
04/22/2020 |
|
42 |
6WEN |
Crystal Structure of ADP ribose phosphatase of NSP3 from SARS-CoV-2 in the apo form |
04/15/2020 |
|
41 |
6YLA |
Crystal structure of the SARS-CoV-2 receptor binding domain in complex with CR3022 Fab [Potent antibody binding to an unexpected highly conserved cryptic epitope of the SARS-CoV-2 Spike] 関連構造 6YM0 |
04/15/2020 |
|
40 |
6YI3 |
The N-terminal RNA-binding domain of the SARS-CoV-2 nucleocapsid phosphoprotein |
04/08/2020 |
|
39 |
6W9Q |
Peptide-bound SARS-CoV-2 Nsp9 RNA-replicase [Identification of a putative peptide-binding site at the dimer interface of the SARS-CoV-2 Nsp9 RNA-replicase, to be published] 6WXD と関連 [一覧 その2] |
04/08/2020 |
|
38 |
6LZG |
Structure of novel coronavirus spike receptor-binding domain complexed with its receptor ACE2 |
03/18/2020 |
|
37 |
7BTF |
SARS-CoV-2 RNA-dependent RNA polymerase in complex with cofactors in reduced condition |
04/08/2020 |
|
36 |
6W9C |
The crystal structure of papain-like protease of SARS CoV-2 |
04/01/2020 |
|
35 |
6M71 |
SARS-Cov-2 RNA-dependent RNA polymerase in complex with cofactors |
04/01/2020 |
|
34 |
|
PanDDA analysis group - 68 structures of COVID-19 protease with various bound fragments (2020-03-11公開分と合わせて75構造) IDリスト: crisp_bio別記事参照 |
03/25/2020 |
|
33 |
6W41 |
Crystal structure of SARS-CoV-2 receptor binding domain in complex with human antibody CR3022 [A highly conserved cryptic epitope in the receptor-binding domains of SARS-CoV-2 and SARS-CoV] |
03/25/2020 |
|
32 |
6W63 |
Structure of COVID-19 main protease bound to potent broad-spectrum non-covalent inhibitor X77 [A taxonomically-driven approach to development of potent, broad-spectrum inhibitors of coronavirus main protease including SARS-CoV-2 (COVID-19)] |
03/25/2020 |
|
31 |
6YB7 |
SARS-CoV-2 main protease with unliganded active site (2019-nCoV, coronavirus disease 2019, COVID-19) [COVID-19 main protease with unliganded active site] |
03/25/2020 |
|
30 |
6W4B |
The crystal structure of Nsp9 RNA binding protein of SARS CoV-2 |
03/18/2020 |
|
29 |
6M0J |
Crystal structure of 2019-nCoV spike receptor-binding domain bound with ACE2 |
03/18/2020 |
|
28 |
6M3M |
Crystal structure of SARS-CoV-2 nucleocapsid protein N-terminal RNA binding domain |
03/18/2020 |
|
27 |
5R7Y |
PanDDA analysis group deposition -- Crystal Structure of COVID-19 main protease in complex with Z45617795 |
03/11/2020 |
|
26 |
5R7Z |
PanDDA analysis group deposition -- Crystal Structure of COVID-19 main protease in complex with Z1220452176 |
03/11/2020 |
|
25 |
5R80 |
PanDDA analysis group deposition -- Crystal Structure of COVID-19 main protease in complex with Z18197050 |
03/11/2020 |
|
24 |
5R81 |
PanDDA analysis group deposition -- Crystal Structure of COVID-19 main protease in complex with Z1367324110 |
03/11/2020 |
|
23 |
5R82 |
PanDDA analysis group deposition -- Crystal Structure of COVID-19 main protease in complex with Z219104216 |
03/11/2020 |
|
22 |
5R83 |
PanDDA analysis group deposition -- Crystal Structure of COVID-19 main protease in complex with Z44592329 |
03/11/2020 |
|
21 |
5R84 |
PanDDA analysis group deposition -- Crystal Structure of COVID-19 main protease in complex with Z31792168 |
03/11/2020 |
|
20 |
6M03 |
The crystal structure of COVID-19 main protease in apo form |
03/11/2020 |
|
19 |
6M17 |
The 2019-nCoV RBD/ACE2-B0AT1 complex |
03/11/2020 |
|
18 |
6M18 |
ACE2-B0AT1 complex |
03/11/2020 |
|
17 |
6M1D |
ACE2-B0AT1 complex, open conformation |
03/11/2020 |
|
16 |
6VXX |
Structure of the SARS-CoV-2 spike glycoprotein (closed state) |
03/11/2020 |
|
15 |
6VYB |
SARS-CoV-2 spike ectodomain structure (open state) |
03/11/2020 |
|
14 |
6VYO |
Crystal structure of RNA binding domain of nucleocapsid phosphoprotein from SARS coronavirus 2 |
03/11/2020 |
|
13 |
6W01 |
The 1.9 A Crystal Structure of NSP15 Endoribonuclease from SARS CoV-2 in the Complex with a Citrate (関連 ID 6X1B 6VWW 6W01 6WLC 6WXC) [Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2.] |
03/11/2020 |
|
12 |
6W02 |
Crystal Structure of ADP ribose phosphatase of NSP3 from SARS CoV-2 in the complex with ADP ribose |
03/11/2020 |
|
11 |
6Y84 |
COVID-19 main protease with unliganded active site (2019-nCoV, coronavirus disease 2019, SARS-CoV-2) |
03/11/2020 |
|
10 |
6VWW |
Crystal Structure of NSP15 Endoribonuclease from SARS CoV-2. (関連 ID 6X1B 6VWW 6W01 6WLC 6WXC) [Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2.] |
03/04/2020 |
|
9 |
6VXS |
Crystal Structure of ADP ribose phosphatase of NSP3 from SARS CoV-2 |
03/04/2020 |
|
8 |
6Y2E |
Crystal structure of the free enzyme of the SARS-CoV-2 (2019-nCoV) main protease |
03/04/2020 |
|
7 |
6Y2F |
Crystal structure (monoclinic form) of the complex resulting from the reaction between SARS-CoV-2 (2019-nCoV) main protease and tert-butyl (1-((S)-1-(((S)-4-(benzylamino)-3,4-dioxo-1-((S)-2-oxopyrrolidin-3-yl)butan-2-yl)amino)-3-cyclopropyl-1-oxopropan-2-yl)-2-oxo-1,2-dihydropyridin-3-yl)carbamate (alpha-ketoamide 13b) |
03/04/2020 |
|
6 |
6Y2G |
Crystal structure (orthorhombic form) of the complex resulting from the reaction between SARS-CoV-2 (2019-nCoV) main protease and tert-butyl (1-((S)-1-(((S)-4-(benzylamino)-3,4-dioxo-1-((S)-2-oxopyrrolidin-3-yl)butan-2-yl)amino)-3-cyclopropyl-1-oxopropan-2-yl)-2-oxo-1,2-dihydropyridin-3-yl)carbamate (alpha-ketoamide 13b) |
03/04/2020 |
|
5 |
6VW1 |
Structure of 2019-nCoV chimeric receptor-binding domain complexed with its receptor human ACE2 |
03/04/2020 |
|
4 |
6LVN |
Structure of the 2019-nCoV HR2 Domain |
02/26/2020 |
|
3 |
6LXT |
Structure of post fusion core of 2019-nCoV S2 subunit |
02/26/2020 |
|
2 |
6VSB |
Prefusion 2019-nCoV spike glycoprotein with a single receptor-binding domain up |
02/26/2020 |
|
1 |
6LU7 |
The crystal structure of COVID-19 main protease in complex with an inhibitor N3 [Structure of Mpro from COVID-19 virus and discovery of its inhibitors: 7BQYと同一論文] |
02/05/2020 |
・日本 PDBjのWebサイトにてID指定、または、https://pdbj.org/mine/summary/ID にて
[例] https://pdbj.org/mine/summary/6VXX
[更新履歴]
2020-05-16 #3の6LXTの参考文献をbioRxiv投稿からCell Research 論文へと更新 (crisp_bio 新型コロナウイルス:スパイクのサブユニットS2の構造と、細胞融合・感染を阻害する脂質化ペプチドの合成)
2020-05-06 ブログの本文記事文字数の上限を超えたため、5月6日PDB更新分から次の記事へ移行: crisp_bio 2020-05-06 新型コロナウイルスのPDB公開構造一覧 - その2
"Structure of Mpro from COVID-19 virus and discovery of its inhibitors" Jin Z, Du X, […] Yang H. Nature 2020-04-09
2020-04-22 4月22日公開の7BV2と7BV1 (bioRxiv 2020-04-10), 6WIQ, および6WJIを追加
2020-04-15 4月8日公開の6W9Qと6YI3, および、15日公開の6YLAと6WENを追加 (いずれも、論文は発表待ち)
2020-04-09 6LU7 (#1)をNature論文とcrisp_bio記事に紐付け
2020-04-09 SARS-CoV-2とACE2複合体のMDシミュレーション動画 "Trajectory 4: A 10 µs simulation of the SARS-CoV-2–ACE2 complex"へのリンクを追加
2020-04-05 The New York Timesが、SARS-CoV-2の全タンパク質の配列データと簡明なアノテーションを提示 "Bad News Wrapped in Protein: Inside te Coronavirus Genome" Corum J, Zimmer C. NYT2020-04-03; How Coronavirus Hijacks Your Cells. NYT 2020-03-13 updated.
2020-04-03 6W41 (#33)にScience論文へのリンクを追加: "A highly conserved cryptic epitope in the receptor-binding domains of SARS-CoV-2 and SARS-CoV" Yuan M, Wu NC [..] Mok CKP, Wilson IA. Science 2020-04-03.
2020-03-31 通し番号修正; Nature 2020-03-30 論文2報へのリンクを追加 (一覧表の#5と#28)
[# 5] "Structural basis of receptor recognition by SARS-CoV-2" Shang J, Ye G, Shi Y, Wan Y [..] Li F. Nature 2020-03-30. [6VW1] Structure of 2019-nCoV chimeric receptor-binding domain complexed with its receptor human ACE2
[#29] "Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor" Lan J, Ge J, Yu J, Shan S[..] Zhang L, Wang X. Nature 2020-03-20. [6M0J] Crystal structure of 2019-nCoV spike receptor-binding domain bound with ACE2
2020-03-31 bioRixv投稿へのリンクを追加 (一覧表の#29) "Crystal structure of the SARS-CoV-2 non-structural protein 9, Nsp9" Littler DR, Gully BS, Colson RN, Rossjohn J. bioRxiv 2020-03-30; 6W4B The crystal structure of Nsp9 RNA binding protein of SARS CoV-2
2020-03-26 03-25新規公開追加の記述修正:新規公開追加 (3件およびPanDDA analsisによるリガンド結合プロテアーゼの構造68件(2020-03-11公開分とあわせて75件)); 論文crisp_bio記事へのリンクを追加し、03-25新規公開については、未発表論文の予定タイトルと思われるテキストを[]内に付記
2020-03-23 PDB登録へのリンク整理; 6LXT (#3)にbioRxiv 2020-03-12投稿と、その紹介にあたるcrisp_bio記事へのリンクを追加
2020-03-18 PDB登録 6M3M 追加
2020-03-12 [更新] 投稿・論文発表があった構造には公開日の部分にcrisp_bio記事または論文へのリンクを埋め込み; PDBj、新型コロナウイルス特集ページ公開 (毎水曜更新) → コチラ
2020-03-12 [注] この一覧は、2002-3年のSARS CoVのPDB登録データを含んでいない ("SARS CoV"検索へのヒットは182件 (2020-03-12時点)。
2020-03-11 [初稿] COVID-19, SARS CoV-2または2019-nCoVにてRCSB PDBを検索した結果25件を一覧に (以後、毎週水曜日に更新されるPDB公開構造と、随時発表される関連論文へのリンクを追加)。
コメント